Dr. Mario A. Fares (Principal Investigator)
Introduction
The
main aim of my research is the understanding of how novel
functions and biological complexity emerge in nature. In
particular, we are interested in identifying the evolutionary
trajectories, at the genome and regulatory levels, to biological
innovations. This aim is relevant not only to the understanding of
species diversification and the emergence of complexity but also
to provide key knowledge for biotechnological and biomedical
developments. Many biological innovations are simply off-limits
for evolution because they involve dramatic changes to organisms
that are often not tolerated by natural selection. However, under
certain conditions, some molecular mechanisms can minimize the
effects of innovative mutations allowing them to survive in the
genome and become eventually fixed, potentially emerging as
adaptive features when the environment changes. Our focus is on
the characterization and use of buffering molecular mechanisms
that provide robustness to mutations allowing the exploration of
larger genotypic networks, and thus originating novel phenotypes.
The main robustness mechanisms we study are heat-shock proteins,
also known as molecular chaperones, and gene and genome
duplications.
To
achieve our goal, we conduct theoretical and experimental research
in a diverse range of fields. In particular, we test specific
evolutionary scenarios by evolving microbes under
laboratory-controlled experiments and analyzing the consequences
of specific evolutionary dynamics at the genome and physiological
levels. A brief description of our specific interests is given
below.
a) The Role of Molecular Chaperones in Regulating Mutational Robustness and Functional Innovation
Heat shock proteins (Hsps) belong to a largely conserved family of proteins that are ubiquitous in living organisms at all levels of complexity, from organelles to microbes and multi-cellular organisms. Hsps perform essential functions in the cell, chief among which is the folding of nascent proteins to acquire their functional conformation. Through the folding of mutated protein versions, some hsps have been suggested to buffer the destabilizing effects of mutations and, in doing so, they allow genomes to explore a wider genotypic network and access otherwise prohibited phenotypes. How do hsps regulate this mutational robustness is unknown. We are mostly interested on the role of GroEL, an essential hsp in prokaryotes and eukaryotic organelles, in regulating mutational robustness and allowing the epistatic interaction between conditionally neutral mutations and innovative destabilizing changes.
b)
The Role of Gene Duplication
in Functional Diversification
Gene duplication is the process by which a gene originates two identical copies, presumably performing identical functions too. Gene duplication can take place at different scales, from single nucleotides to entire genomes. While the classic theory of Susumo Ohno established that after duplication one gene copy, devoid of selection pressures, can explore novel functions alternative to the ancestral ones, evolution by gene duplication seems more complex than predicted by theory: both the genomic background in which it arises and the mechanism of duplication having significant impact on determining the functional fates of duplicates. What seems less doubtful is that major organismal diversification events are concomitant with gene and genome duplications. However, it remains undefined the mechanism through which gene duplication leads to innovations. In my group, we investigate mutational robustness mediated by gene duplication as a starting point in the re-wiring of the epistatic and regulatory relationships between genoytpes, a cause prima facie of innovations.
c) Molecular Coevolution
Coevolution
is
classically defined as the reciprocal natural selection between
two interacting populations. From the molecular perspective,
coevolution can be readily understood as the reciprocal selective
constraints between two interacting molecules. Within the
framework of molecular coevolution I include also epistasis
because certain genomic background can facilitate previously
prohibited mutations that, in turn, modify the epistatic potential
of the new genomic context. We are interested in those signatures
of coevolution that are the result of molecular coadaptation. For
example, physically interacting proteins exercise reciprocal
selection upon one another that is translated into a coadaptation
process which leaves coevolution signatures in the amino acid
sequences. Coevolution has many theoretical and pragmatic
application, probably the most sound being the in silico
identification of protein-protein physical interactions or the
prediction of three-dimensional protein folds. In my group, we
develop mathematical models and computational tools to identify
molecular coevolution within and between protein sequences. We are
particularly interested in identifying coadaptation dynamics in
general, protein-protein interactions and functional shifts
leading to protein functional promiscuity.
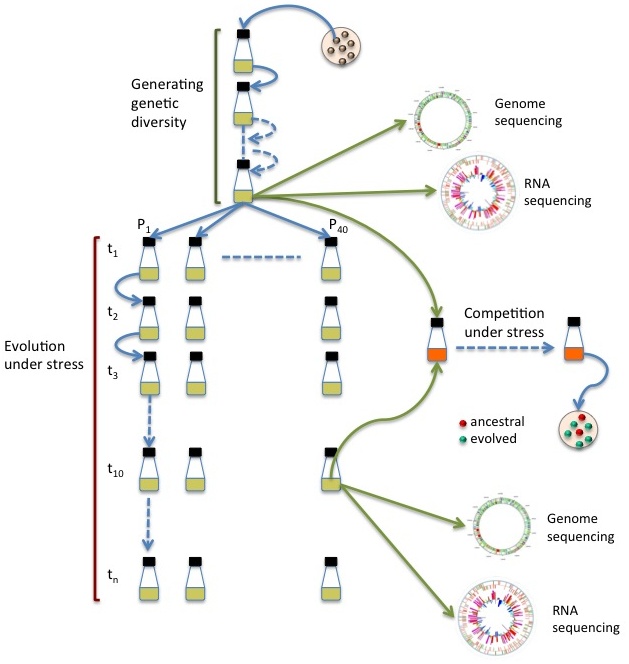
d) Experimental Evolution





 Fitness Landscape and Adaptive traps
Fitness Landscape and Adaptive traps